Intro to GitHub with RStudio and HPC systems
Published:
GitHub is one of those things that many people in research have vaguely heard of, know they probably should be using, and then avoid for far too long. That was definitely me at first.
If you are not building apps, working in a large software team, or spending all day in the command line, GitHub might not feel like its particularly made for you. A lot of the explanations online are aimed at developers, which can make it feel less relevant than it actually is for academic workflows.
But if you write code for analyses, make figures in R, run jobs on an HPC, or want your work to be easier to organise, reproduce, and share, hopefully I can convince you GitHub is very helpful and that the earlier you start using GitHub in a project the better. The setup and basic day-to-day workflow is also much simpler than it first appears.
In this post, I want to go through how I actually use GitHub in research:
- why I use it
- key terms
- setting up in RStudio
- my day-to-day in RStudio,
- using Git with HPC systems
Why I use GitHub
For me, GitHub is useful for three main reasons: change tracking, collaboration, and project organisation.
1. Change tracking
![]()
GitHub keeps a history of changes to your files. Instead of saving endless versions of the same script with names like analysis_final.R analysis_final_v2.R and analysis_final_FINAL.R you can keep one script under the same name, and the changes you make to the script is saved as commits with messages explaining what changed. If at any point you mess up your code and you dont know why- you can recover your file to a previous commit.
If you are anything like me, that means you can stop cluttering code with old fragments you are too nervous to delete “just in case” and you do not have to keep an very long confusing record of backup files across your desktop.
2. Collaboration
GitHub is set up for collaboration and there are many ways I find it useful even if I am not part of a massive team building fancy apps or software.
First I have added my supervisors to my past repositories, making it easier to discuss issues, key changes or questions with code.
Next, if you are developing code with others, multiple people can work on the same project without emailing scripts back and forth. This also avoids keeping versions in shared folders with the fear someone might delete or overwrite something. GitHub is especially useful when someone else needs to check code, fix an error, or run something on a different system.
I have also found the collaboration setups really useful for myself moving cleanly between computers and systems. For example me-on-my-laptop to me-on-the-HPC or to me-on-a-different-computer.
Finally, once you are done with a project and you have published it (!), having you code already on GitHub is a huge plus. You then don’t need to worry that in transfering files to GitHub for open science all your code will break- it won’t because it its already on there!
3. Better project organisation
While not innately a specific benefit, by using GitHub I have been influenced to have better file management in each project. GitHub works in the way where each project is kept within one neat, self-contained repository. Therefore, all my projects now start with the same initail structure, making my projects easier to understand, even when I have put is down for a while.
So to sum up, I view using GitHub as a way to keep everything together in a neat, transferable, reproducible package. Great for day-to-day and will make your/our lives easier in the long run.
There is just that pesky initial learning curve…
Key terms
GitHub, like everything else in life, comes with some jargon. The good news is that you do not need much of it to get started.
Hoepfully this will clear some things up!
| Term | What it means | How I think about it |
|---|---|---|
| Git vs GitHub | Git is the version control system (tracks changes). GitHub is the online platform (hosts and shares projects). | Like R vs RStudio — Git is the engine, GitHub is the interface. |
| Repository (repo) | A project folder tracked by Git. In R I set this as my “Working Directory” | I treat each repo as one body of work (paper, chapter, project). |
| File paths | The way code locates files in your computer/repo. | e.g ./code/data_import.R
|
| Stage | The step where you tell Git which files you want to include in the next save point. | These are the files I want to upload to GitHub” |
| Commit | A saved checkpoint in your project history, with a message describing the change. | A labelled save point. Good messages means your future self will thank you! |
| Branches & merging | Parallel versions of code that can later be combined. | Useful for teams. I don’t really use this day-to-day (yet). |
One important limitation
GitHub is for code and associated project files, not for storing huge raw datasets.
GitHub does not store files over 2 GB , trying to do so will throw an error so large data files should be stored elsewhere and linked or documented properly instead- an example of how I do this below!
Setting up GitHub with a new RStudio project

When starting from complete scratch (eg on a brand new computer), my general setup process is:
- Install Git on the computer -> and tell RStudio where to locate that Git program
- Create a new repository on GitHub
- Create or connect an R project in RStudio
- Link the local project to the repository
In practice, (as I already have installed git on my computer) I often do it in this order:
Step 1: Make the repository on GitHub
I create an empty repository online, usually with:
- a repository name
- private visibility to begin with
- a README file
Step 2: Clone it into RStudio
In RStudio, you can create a new .RProj from File>New Project>Version Control>Git>Input repository URL (often https://github.com/yougithubusername/yourreponame.git.)
That downloads the online repository to your computer and creates the linked R project locally.
Step 3: Populate the file structure
Once the repo exists locally, I like to set up my usual file structure, with the knowledge I then have to leave that structure alone so less chance of the code breaking.
Using R Projects changed my life, because it sets the working directory automatically, keeps code and data together, and can link with Git and GitHub. Previously I have code files and data files saved accross my computer and was always losing things or breaking my code when the file route changed.
One slightly confusing thing at first is that empty folders do not show up on GitHub. So if you create a nice folder structure locally and then panic because GitHub is not showing it, that is normal. Once files exist inside those folders and you commit them, they will appear.
A my usual structure looks something like this:
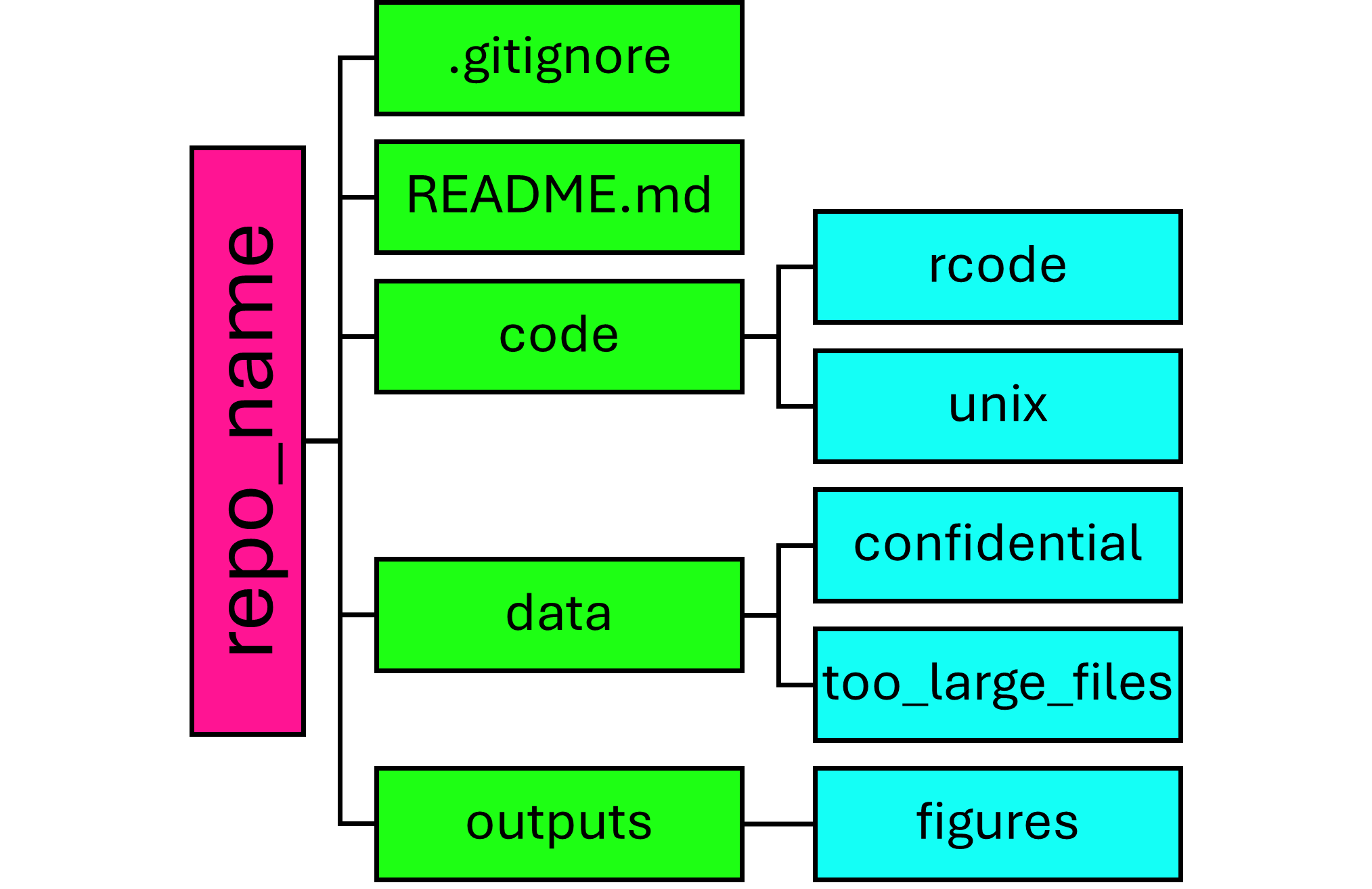
Assuming you have already made the RProj, and linked with GitHub, so already have the empty repo made with the REDAME and .gitignore - this can save you some time making subfolders ![]()
# create folder structure
dir.create(c(
"code/rcode",
"code/unix",
"data/confidential",
"data/too_large_files",
"outputs/figures"
), recursive = TRUE, showWarnings = FALSE)
I have settled on this structure for a few reasons.
First, it keeps file paths tidy and portable. If everything lives inside one project folder, it is much easier to move the project between locations or machines without breaking links to data and outputs.
Second, it makes easy to seperate what files should and should not go to GitHub. I can just set the .gitignore up to ignore certain subfolders and then forget about it.
Using .gitignore
The .gitignore file tells Git which files or folders to ignore. You essentially open the .gitignore file and write down the file paths you want to ignore.
In my projects, I use it mainly for:
- data files that are too large for GitHub (>2GB)
- confidential files that should never be uploaded
- occasional temporary or system-specific files
So for me I write in the .gitignore file something like:
data/too_large_files/
data/confidential/
*This must be pushed to GitHub to function I believe.
Using README.md
I also strongly recommend having a README.md file in each repository. If you make a repository public, or share with collaborators or have to put the project down for a while, the README is the landing page where you explain what the project is, how the code is organised, what order scripts should be run in, and where any related publication can be found.
My day-to-day Git workflow in RStudio
Once a project is set up, my actual Git workflow is very simple.
After I have done some coding and saved my changes locally, I usually go into the terminal in RStudio and run:
git status
git add code/
git commit -m "brief description of what changed"
git push origin main
That is basically it.
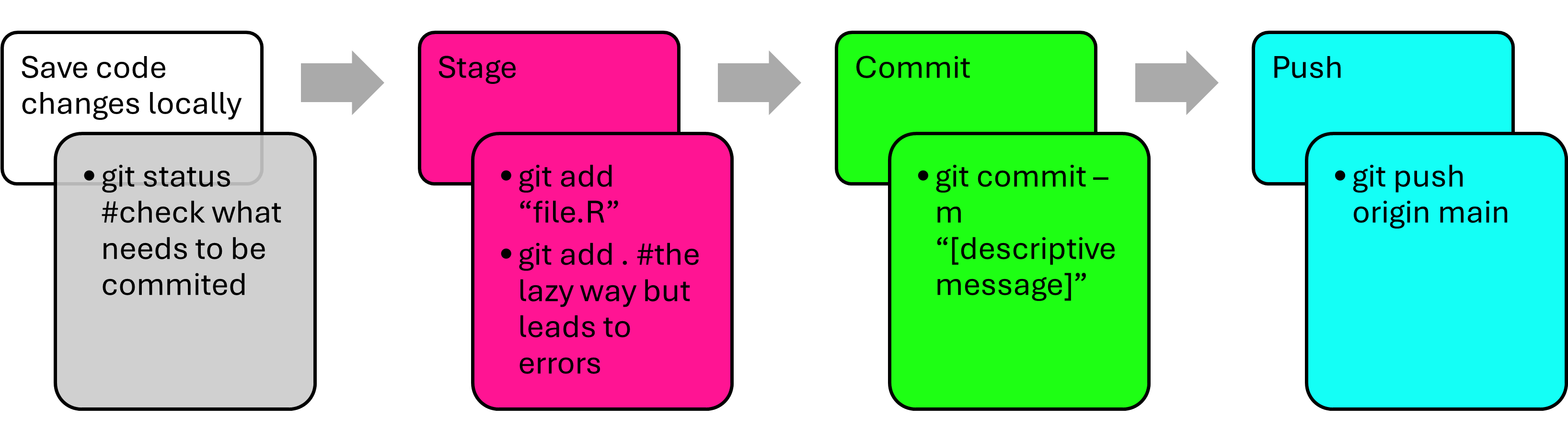
Here is what each command is doing:
git status
This shows what is different between your local project and the version on GitHub. It also shows you the name of your branch- if you havent branched off at all this will usually be “main”.
It is my first step every time because it tells me what has changed and what Git is seeing.
git add
This stages the files I actually want to include in the next commit.
For example, if I only changed code files, I can add just the code directory:
git add code/
That is much safer than blindly adding everything.
I used to always use:
git add .
The . means “the current directory”, so this stages everything Git detects as changed in that location.
This can be useful, but only when you are very sure what has changed. If you have large files, confidential files, or random clutter in a project, git add . is a great way to get errors from uploading something over 2GB, or to stage something you did not mean to upload.
So my general advice is:
- use targeted
git addcommands when possible - use
git statusfirst to identify large randomly generated clutter - only use
git add .when you are confident about what is new in the repo
If like me you have accidentally tried to push something too large to GitHub-and have found out by recieving an error- this might be the code to help. I have not used it in a while because after the faff of getting out of that error I have been very carefull to avoid this. You will need to write it in the terminal in R.
git reset HEAD~1
git rm --cached path/to/large_file
git commit -m "remove large file"
git push
Its a faff because if you have pushed everything you might not know exactly what your large file is- then you would need to uncache the whole push or something? This is a mistake everyone does at least once and tries to never again!
git commit -m " "
This saves a checkpoint in the project history and attaches a message to it.
For example:
git commit -m "clean up SDM workflow and remove unused script"
Lazy me writes “.” as a message, however I learnt that lesson when I wanted to look back at my code history after I made a specific faulty change and just saw a bunch of commits called ".". Its like…not labelling a sample well in the lab- you can find info out by looking at dates etc but its much more useful having a intuative name or well labelled commit!
git push origin main
This uploads those committed changes to GitHub.
If I have just been working on my own machine and want the online version updated, this is the final step.
git pull origin main
This does the opposite of git push — it downloads newer commits from GitHub onto your local machine.
If you are working on a single machine on your own, you may not use this much. It becomes important when you are:
- you have made changes on another machine
- working across multiple computers (e.g. laptop and HPC)
- collaborating with other people
In those cases, git pull makes sure you are working with the most up-to-date version of the project. git log will also show you the history of commits on the project.
For further reading on commands for git- see the GitHub cheatsheet.
Why I prefer the terminal over the Git tab
RStudio does have a Git tab, and you absolutely can use it.
I do not. I find the terminal much clearer and faster once you get used to it. It feels more direct, and it makes the underlying steps more obvious. I think of the Git tab as about as useful as manually clicking dropdowns to import csv files into R- using the clicking method is intuative for learning but doesnt get you far in the long run.
How I use Git/GitHub on an HPC
When doing bioinformatics, I often work partly on my own computer and partly remotely on an HPC. The HPC is a powerful computer cluster used for high-memory tasks such as aligning genomic reads, with users interacting through the command line and submitting scripts.
When doing HPC work, you quickly find its easier to write new scripts locally on a nice editing software (eg. Notepad ++), but sometimes for problem solving you make little changes within the HPC system. Therefore you generally end up with two slightly different files on your local PC and the HPC that you dont know which is the working/most up to date script.
Cue ![]() GitHub
GitHub ![]()
My HPC bioinformatics workflow looks like this:
- Edit code locally (in Notepad ++)
- Commit and push changes from RStudio terminal (because I have already linked my GitHub repo to my RProj and I am always tickering in there anyway!)
- Pull those changes on the HPC
- Run the job there
For example, locally in RStudio I might run: 
git status
git add code/unix/
git commit -m "update VCF output script"
git push origin main
Then on the HPC I run:
git status
git pull origin main
and the updated scripts are there. If I make a quick edit in the HPC I do it the other was round- push from the HPC and pull from my local PC.
Setting up GitHub authentication on an HPC
To connect an HPC to GitHub, you usually need to set up SSH authentication.
- log in to the HPC
- generate an SSH key
ssh-keygen -t ed25519 -C "your_email@example.com" copy the public key into your online GitHub settings
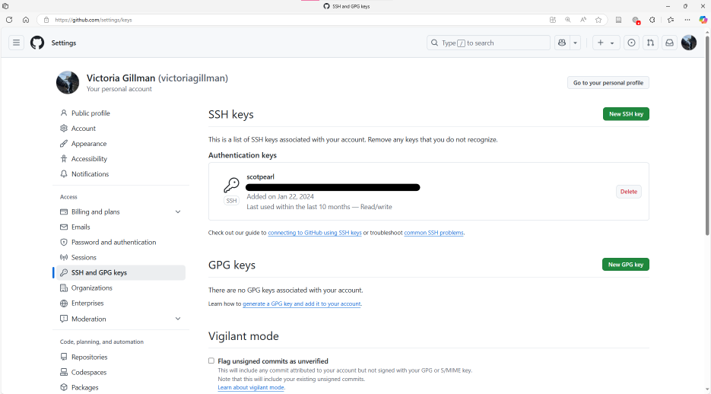
- test the connection
ssh -T git@github.com - clone your repository onto the HPC
git clone git@github.com:username/repositoryname.git
You only have to do this once, then afterwards it tends to just work quietly in the background.
One final GitHub tip
At one point, I noticed my commits were not being attributed properly to my GitHub profile.
Git was using a system default username and email rather than the ones linked to my account. To fix this you need to set your global Git username and email in the terminal so commits are recognised correctly
That looks like this:
git config --global user.name "Your Name"
git config --global user.email "your.email@example.com"
This was mostly important because I like the little contribution heatmap on GitHub
Final thoughts
The main thing I want to get across is that GitHub does not have to mean becoming a software engineer.
For a lot of academic work, especially if you mostly code alone, GitHub is actually very simple and useful:
- keep each project in a clean folder structure
- use GitHub to track code changes
- write sensible commit messages
- keep large or confidential files out of the repo
- use the same repository across your local machine and HPC
If you are new to GitHub, once the initial setup and learning curve is done, the everyday workflow often comes down to four commands: status, add, commit, and push

Helpful resources
If you want to go a bit further, two resources I recommend are:
I also have a presentation version of this, please to get in touch if that would be useful for your group. ![]()
